SMIDs: Unique identifiers for biogenic small molecules in C. elegans.
1. The issue
Small molecules/secondary metabolites are referred to by a plethora of names and abbreviations. Some compounds are referred to by more than 10 different names, and in certain cases different compounds are referred to by the same name. Significantly, there is no established system for naming newly identified metabolites that would permit database searching for small molecule metabolites in the same manner as for genes.Example I highlights the large number of acceptable names for a signaling molecule recently identified from C. elegans.
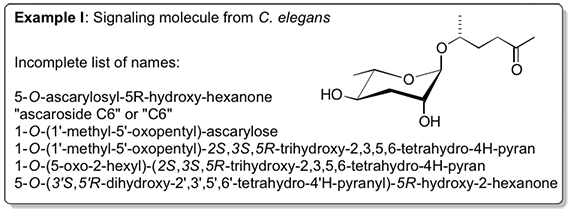
Ambiguities and parallel usage have prevented the development of effective text mining tools for small molecules. As a result, effective sharing of small-molecule data in chemical biology and metabolomics is virtually impossible. Even experienced researchers familiar with the chemical nomenclature often have difficulty locating references for a specific substance. Researchers that are less knowledgeable with chemical nomenclature face even greater difficulties when trying to locate a specific substance or reference.
2. Existing naming schemes do not offer a viable solution
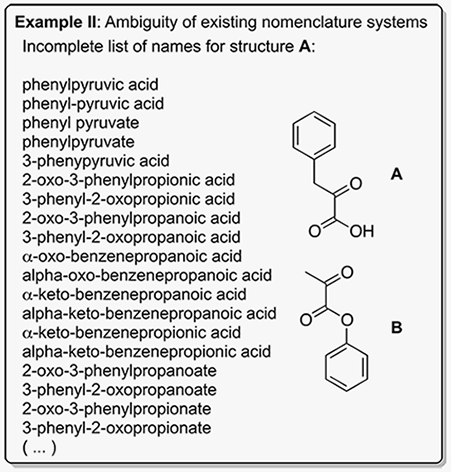
CAS: The Chemical Abstracting Service (CAS) assigns every new compound presented in the literature a unique Chemical Abstracts registry number (CAS#). For example, using the CAS system the compounds shown above in Example I and II are referred to as 946524-24-9 (Example I) 156-06-9 (Example II, Structure A), and 2149-49-7 (Example II, Structure B). Although the CAS system is useful for archiving the chemical literature, CAS numbers are cumbersome to use in scientific writing as they have no recognition value. Importantly, many biological journals are not indexed by CAS.
IUPAC: The IUPAC nomenclature system is highly sophisticated. As a result, derivation and interpretation of IUPAC names requires extensive chemical knowledge. Non-chemists are frequently unable to determine whether two IUPAC names refer to the same compound or not. In addition, IUPAC names are often exceedingly long and complicated and thus unsuitable for use in scientific writing.
SMILES: SMILES are useful as technical, unambiguous descriptors of chemical structures, but unsuitable as in-text identifiers.
3. A new proposal for naming biogenic small molecules/secondary metabolites in C. elegans: SMIDs
I. Small molecules newly identified from the nematode C. elegans are assigned a unique biogenic Small Molecule Identifier (SMID) consisting of four lower case non- italicized letters that refer to the general structural class of the compound, followed by a pound sign and a number. This scheme is comparable to that used for genes and proteins: daf-22 (three letters, italicized, lower case) or DAF-22 (non-italicized, upper case).
Examples:Many pheromones in C. elegans belong to a class of glycosides known as ascarosides. Therefore, ascr was chosen as the four-letter SMID for this class of compounds.
- ascr#1:"daumone" or "C7" or "(6R-(tetrahydro-3'R,5'R-dihydroxy-6'S-methyl-2Hpyran-2'R-yloxy)-heptanoic acid"
- ascr#4:"nematone-1" or "5R-(3'-O-beta-D-glucosyl-tetrahydro-3'R,5'R-dihydroxy-6'Smethyl-2H-pyran-2'R-yloxy)-2-hexanone"
Similarly, steroids called dafachronic acids that regulate C. elegans development have been assigned the four-letter SMID "dafa":
- dafa#1:("delta4-dafachronic acid" or "3-keto-4-cholestenoic acid")
- dafa#2:("delta7-dafachronic acid" or "3-keto-7,(5a)-cholestenoic acid")
II. Stereoisomers are distinguished by the addition of a second numeral. The first discovered stereoisomer of any compound will be named with the ending .1, i.e. xxxx#x.1. For example, in the case of ascr#6, (-)-5R-(3'R,5'R-dihydroxy-6'S-methyl-(2H)-tetrahydropyran-2'-yloxy)-2R-hexanol would be ascr#6.1, and (-)-5R-(3'R,5'R-dihydroxy-6'S-methyl -(2H)-tetrahydropyran-2'-yloxy)-2S-hexanol would be ascr#6.2.
III. The SMID database is maintained by Tyler Schwertfeger of the Schroeder Lab at the Boyce Thompson Institute, with database support by the Mueller lab at the BTI. Previously, Joshua Judkins and Maro Kariya (both of Boyce Thompson Institute and Cornell University) have curated the database, in collaboration with Wormbase. For each C. elegans metabolite, SMID-DB.org provides::
- Structure (structural drawing, SMILES)
- Compound ID (common names, CAS, Beilstein, IUPAC)
- Original reference(s)
- List of references that mention the compound
- Genes in associated pathways (e.g. receptors, biosynthetic enzymes)
All gene entries at SMID-DB.org are linked to Wormbase.org.
For questions and comments or to submit new compounds, please contact smid-db@cornell.edu.